|
I have a Ph.D. and a masters degree in biophysics, and a 5-year degree in physics. Before joining the Foqum team, I was a postdoc researcher at the IT University of Copenhagen, at the Robotics, Evolution and Art Lab, with Sebastian Risi.
I am interested in problems related to biology and life, but my tools and perspective are those of a physicist. I enjoy developing simple models of complex systems and exploring them through computer simulations.
My most recent academic work is on artificial neural networks, an amazing meeting point of disciplines, which includes biology, medicine, physics, robotics, computer science and more.
The evolution of complex neural networks still faces unsolved problems. For example, user-guided evolution is very fragile, and learning new tasks is challenging, if at all possible. Mixed-initiative has difficulties of its own. For instance, humans tend to get lost trying to develop a useful intuition relating network structure and function.
We use modular structures to address such problems. Different modules can isolate—and protect—functional sub-networks, and the resulting network has a higher level structure that is more familiar to humans.
We are also intrigued by the possibility of introducing evolutionary techniques and the human element in deep learning techniques.
Sounds interesting, tell me more!
Thesis Project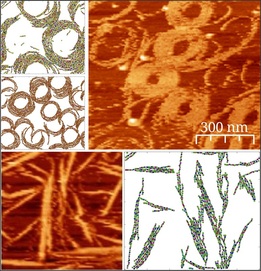
During my PhD project I have been studying the dynamics of FtsZ filaments. FtsZ is a protein found in bacteria (and also in archaea) and it plays an essential role in cell division through the formation of the Z-ring, which is involved in the production of the constriction forces needed to split the cell in two. The details of this important process are still quite far from being understood.
We used Montecarlo simulations of FtsZ monomers on a fine-grained lattice (this means that the proteins occupy several lattice points, allowing for subtle movement) and compared the results with atomic force microscopy images of FtsZ on a flat surface. Our aim was to identify the essential interactions among the proteins that result in the very dynamic aggregates that are found in the experiments. In this way, I hope to have shed some light into the much more complex cellular environment, where experiments are more difficult to interpret.
Do you want to know a bit more about my research with FtsZ?
At the end of my stay I developed, with the help of Juan Carlos González-Avella, a Java applet that could create different types of networks with many adjustable parameters and run voter dynamics on them, showing some statistics of the system during the simulation. Sadly, Java applets are a safety risk and it was recently removed from IFISC's website.
New! I have uploaded the model to NetLogo Modeling Commons, where you can download it or run it in the web version of NetLogo (under development, and not displaying the model correctly. I will try to create a new version that works better there). I still need to add information and the model description (which I can't because of some technical issues).
|